Methods for zeroinfl Objects
predict.zeroinfl.RdMethods for extracting information from fitted zero-inflated
regression model objects of class "zeroinfl".
Usage
# S3 method for class 'zeroinfl'
predict(object, newdata,
type = c("response", "prob", "count", "zero"), na.action = na.pass,
at = NULL, ...)
# S3 method for class 'zeroinfl'
residuals(object, type = c("pearson", "response"), ...)
# S3 method for class 'zeroinfl'
coef(object, model = c("full", "count", "zero"), ...)
# S3 method for class 'zeroinfl'
vcov(object, model = c("full", "count", "zero"), ...)
# S3 method for class 'zeroinfl'
terms(x, model = c("count", "zero"), ...)
# S3 method for class 'zeroinfl'
model.matrix(object, model = c("count", "zero"), ...)Arguments
- object, x
an object of class
"zeroinfl"as returned byzeroinfl.- newdata
optionally, a data frame in which to look for variables with which to predict. If omitted, the original observations are used.
- type
character specifying the type of predictions or residuals, respectively. For details see below.
- na.action
function determining what should be done with missing values in
newdata. The default is to predictNA.- at
optionally, if
type = "prob", a numeric vector at which the probabilities are evaluated. By default0:max(y)is used whereyis the original observed response.- model
character specifying for which component of the model the terms or model matrix should be extracted.
- ...
currently not used.
Details
A set of standard extractor functions for fitted model objects is available for
objects of class "zeroinfl", including methods to the generic functions
print and summary which print the estimated
coefficients along with some further information. The summary in particular
supplies partial Wald tests based on the coefficients and the covariance matrix
(estimated from the Hessian in the numerical optimization of the log-likelihood).
As usual, the summary method returns an object of class "summary.zeroinfl"
containing the relevant summary statistics which can subsequently be printed
using the associated print method.
The methods for coef and vcov by default
return a single vector of coefficients and their associated covariance matrix,
respectively, i.e., all coefficients are concatenated. By setting the model
argument, the estimates for the corresponding model components can be extracted.
Both the fitted and predict methods can
compute fitted responses. The latter additionally provides the predicted density
(i.e., probabilities for the observed counts), the predicted mean from the count
component (without zero inflation) and the predicted probability for the zero
component. The residuals method can compute
raw residuals (observed - fitted) and Pearson residuals (raw residuals scaled by
square root of variance function).
The terms and model.matrix extractors can
be used to extract the relevant information for either component of the model.
A logLik method is provided, hence AIC
can be called to compute information criteria.
Examples
data("bioChemists", package = "ModTools")
fm_zip <- zeroinfl(art ~ ., data = bioChemists)
plot(residuals(fm_zip) ~ fitted(fm_zip))
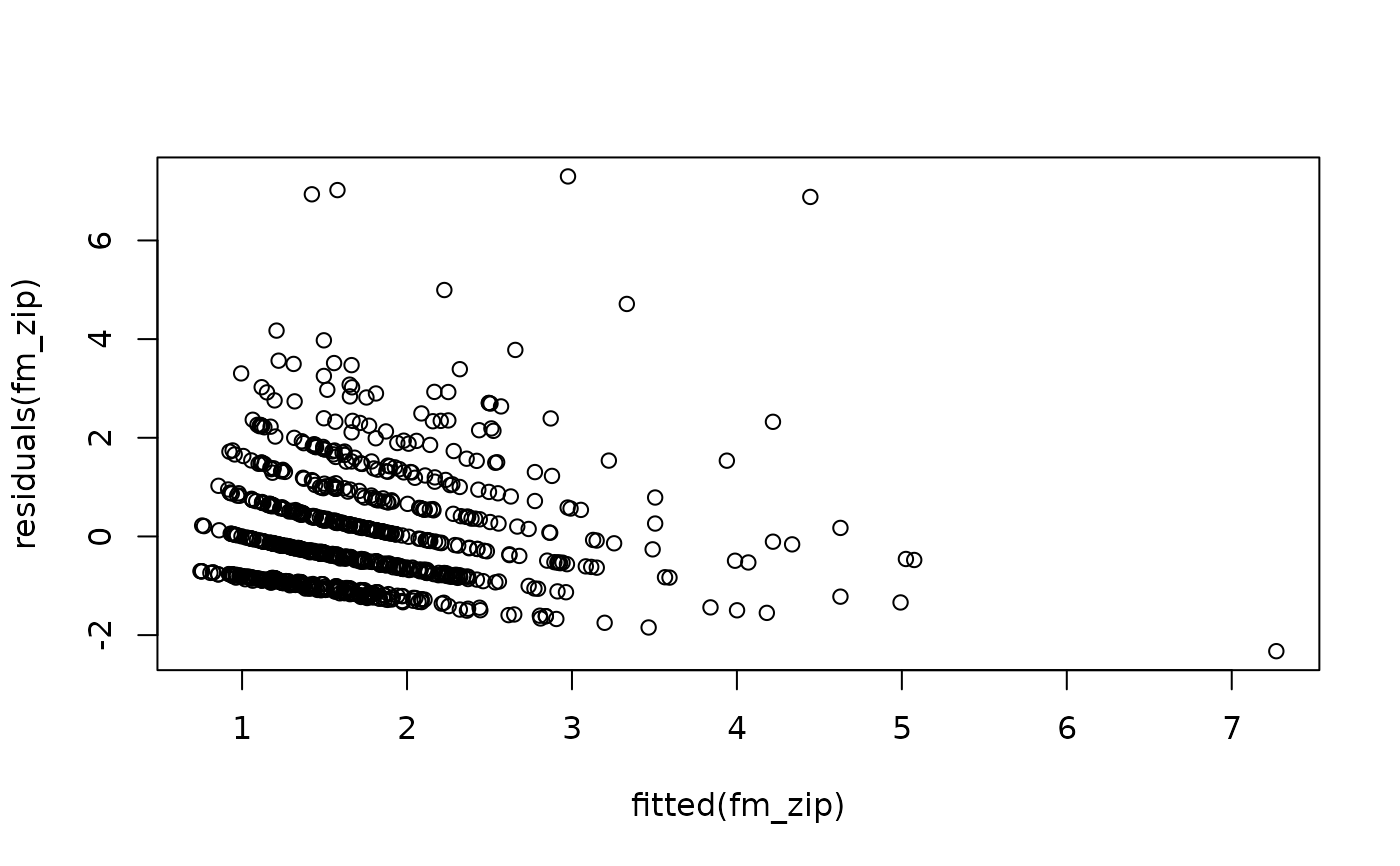 coef(fm_zip)
#> count_(Intercept) count_femWomen count_marMarried count_kid5
#> 0.640838981 -0.209144354 0.103750213 -0.143320239
#> count_phd count_ment zero_(Intercept) zero_femWomen
#> -0.006166143 0.018097723 -0.577060250 0.109751512
#> zero_marMarried zero_kid5 zero_phd zero_ment
#> -0.354017556 0.217095103 0.001274820 -0.134114310
coef(fm_zip, model = "count")
#> (Intercept) femWomen marMarried kid5 phd ment
#> 0.640838981 -0.209144354 0.103750213 -0.143320239 -0.006166143 0.018097723
summary(fm_zip)
#>
#> Call:
#> zeroinfl(formula = art ~ ., data = bioChemists)
#>
#> Pearson residuals:
#> Min 1Q Median 3Q Max
#> -2.3253 -0.8652 -0.2826 0.5404 7.2976
#>
#> Count model coefficients (poisson with log link):
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) 0.640839 0.121307 5.283 1.27e-07 ***
#> femWomen -0.209144 0.063405 -3.299 0.000972 ***
#> marMarried 0.103750 0.071111 1.459 0.144567
#> kid5 -0.143320 0.047429 -3.022 0.002513 **
#> phd -0.006166 0.031008 -0.199 0.842376
#> ment 0.018098 0.002294 7.888 3.07e-15 ***
#>
#> Zero-inflation model coefficients (binomial with logit link):
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) -0.577060 0.509386 -1.133 0.25728
#> femWomen 0.109752 0.280082 0.392 0.69517
#> marMarried -0.354018 0.317611 -1.115 0.26501
#> kid5 0.217095 0.196483 1.105 0.26920
#> phd 0.001275 0.145263 0.009 0.99300
#> ment -0.134114 0.045243 -2.964 0.00303 **
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> Number of iterations in BFGS optimization: 19
#> Log-likelihood: -1605 on 12 Df
logLik(fm_zip)
#> 'log Lik.' -1604.773 (df=12)
coef(fm_zip)
#> count_(Intercept) count_femWomen count_marMarried count_kid5
#> 0.640838981 -0.209144354 0.103750213 -0.143320239
#> count_phd count_ment zero_(Intercept) zero_femWomen
#> -0.006166143 0.018097723 -0.577060250 0.109751512
#> zero_marMarried zero_kid5 zero_phd zero_ment
#> -0.354017556 0.217095103 0.001274820 -0.134114310
coef(fm_zip, model = "count")
#> (Intercept) femWomen marMarried kid5 phd ment
#> 0.640838981 -0.209144354 0.103750213 -0.143320239 -0.006166143 0.018097723
summary(fm_zip)
#>
#> Call:
#> zeroinfl(formula = art ~ ., data = bioChemists)
#>
#> Pearson residuals:
#> Min 1Q Median 3Q Max
#> -2.3253 -0.8652 -0.2826 0.5404 7.2976
#>
#> Count model coefficients (poisson with log link):
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) 0.640839 0.121307 5.283 1.27e-07 ***
#> femWomen -0.209144 0.063405 -3.299 0.000972 ***
#> marMarried 0.103750 0.071111 1.459 0.144567
#> kid5 -0.143320 0.047429 -3.022 0.002513 **
#> phd -0.006166 0.031008 -0.199 0.842376
#> ment 0.018098 0.002294 7.888 3.07e-15 ***
#>
#> Zero-inflation model coefficients (binomial with logit link):
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) -0.577060 0.509386 -1.133 0.25728
#> femWomen 0.109752 0.280082 0.392 0.69517
#> marMarried -0.354018 0.317611 -1.115 0.26501
#> kid5 0.217095 0.196483 1.105 0.26920
#> phd 0.001275 0.145263 0.009 0.99300
#> ment -0.134114 0.045243 -2.964 0.00303 **
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> Number of iterations in BFGS optimization: 19
#> Log-likelihood: -1605 on 12 Df
logLik(fm_zip)
#> 'log Lik.' -1604.773 (df=12)